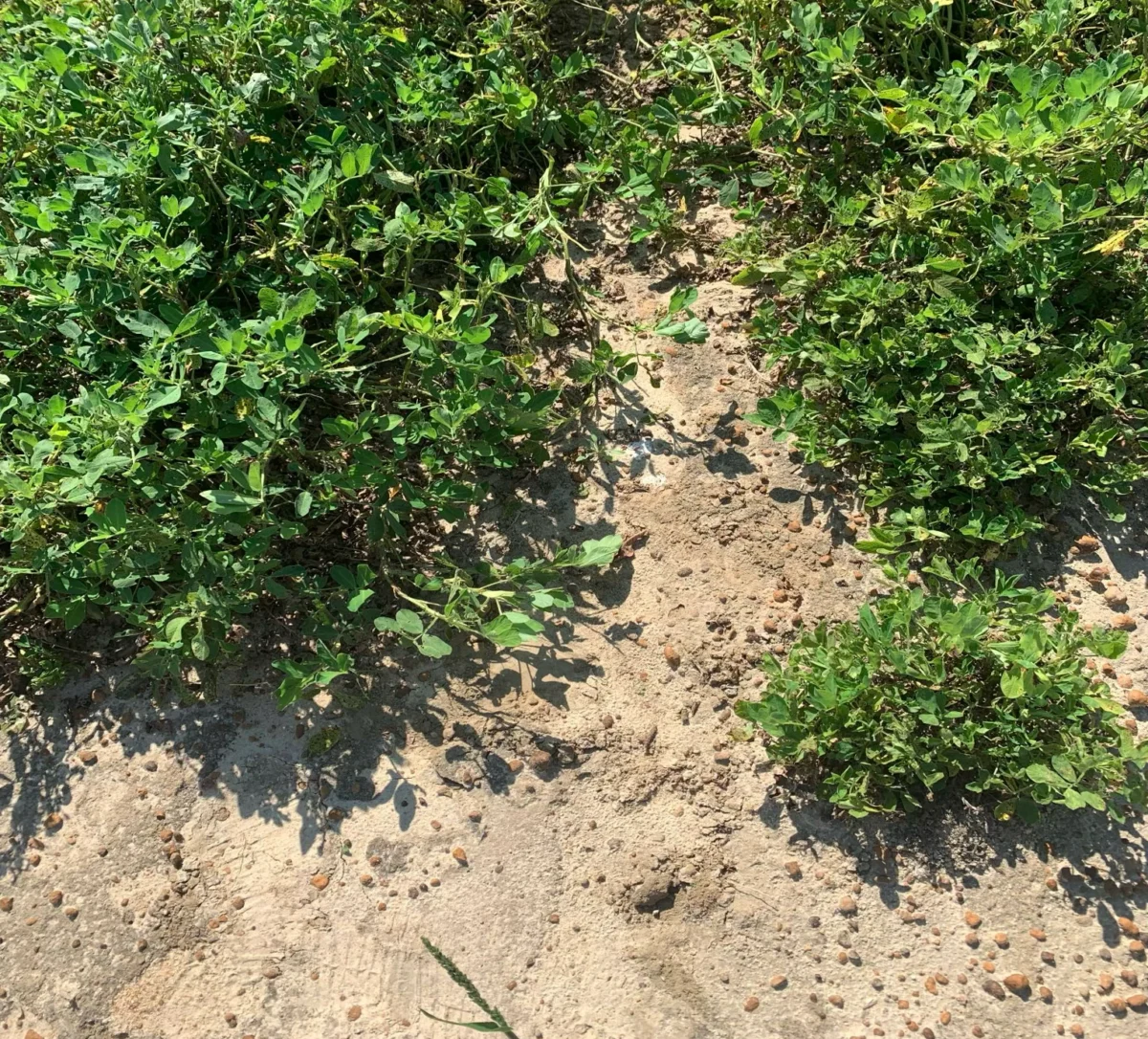
For two decades, a silent adversary has plagued agricultural yields across the globe. Tomato Spotted Wilt Virus (TSWV) has been responsible for billions of dollars in losses, leaving farmers and plant breeders in a protracted, often losing, battle. Despite the urgency, the genetic mechanism behind resistance remained elusive, shielded by the limitations of traditional genomic tools.
That changed with the development of "Khufu," a pioneering genomic methodology developed by the HudsonAlpha Institute for Biotechnology. By moving away from the rigid constraints of a single reference genome and embracing the fluidity of pangenomics, researchers have finally cracked the code of TSWV resistance. This breakthrough not only provides a definitive answer to a decades-old mystery but heralds a new era in precision breeding.
Main Facts: The Khufu Paradigm Shift
At its core, the Khufu approach is designed to maximize the utility of low-pass, short-read whole genome sequencing. Historically, short-read sequencing has been criticized for its inability to capture complex structural variations, often forcing researchers to align sequences against a "linear" reference genome. This process frequently introduces "reference bias," where unique or complex genetic variations—the very ones that often dictate resistance—are discarded or misaligned.
Khufu, paired with its specialized add-on package, KhufuPAN, circumvents this by generating custom pangenome graphs. Instead of comparing an individual plant to a singular, static sequence, Khufu maps genomic data within a broader context that mirrors actual population diversity.
The transformative result? The discovery that TSWV resistance was not governed by a single nucleotide polymorphism (SNP), as many had long hypothesized, but by a sophisticated structural variant: a duplicated gene cassette containing four copies of a glutamate receptor gene. This copy number variation (CNV) acts as a genetic "dosage" mechanism—the more copies a plant carries, the stronger its resistance to the virus.
Chronology: Two Decades of Scientific Stagnation and Recent Breakthrough
The journey to this discovery is a testament to the evolution of genomic technology.
- The 2000s – Early 2020s: For twenty years, breeders attempted to map the TSWV resistance locus using traditional marker-assisted selection. These methods relied heavily on SNP-based arrays and linear reference alignment. While these tools were effective for simple traits, they consistently failed to capture the structural complexity of the TSWV locus, leading to inconsistent phenotypic associations and stalled breeding programs.
- The Development Phase: Recognizing that the "missing heritability" was likely hidden in structural variations, the team at HudsonAlpha developed the Khufu pipeline. The goal was to create a cost-effective method to scale sequencing across thousands of individuals while maintaining high-resolution mapping.
- The Implementation: Using a large-scale segregating population, the team applied the KhufuPAN framework. By analyzing the structural landscape of the population, the researchers identified the glutamate receptor gene duplication.
- The Validation: The team correlated the number of gene copies with disease outcomes. Plants with four copies showed high resistance, those with two or three showed moderate resistance, and those with zero copies were entirely susceptible to the virus.
- Present Day: The methodology has transitioned from a research curiosity into a standardized tool for breeders, allowing for the direct selection of optimal copy number configurations rather than relying on variable field pressure tests.
Supporting Data: Decoding the Glutamate Receptor Cassette
The data generated by the Khufu approach offers a compelling narrative of genetic dosage. In traditional genomics, a researcher might look for a mutation in a single gene. Here, the "mutation" is the expansion of a genetic block.
The structural variation identified involves a tandem duplication. By utilizing the KhufuPAN graph, researchers could visualize the "paths" through the genome that included these four distinct copies. Statistical analysis of the population showed a clear linear correlation:
- Full Resistance (4+ copies): Individuals exhibiting this genotype showed negligible signs of TSWV infection even under high-pressure exposure.
- Moderate Resistance (1-3 copies): These plants displayed delayed symptom onset and lower viral titers, providing a buffer but not total immunity.
- Susceptibility (0 copies): In the absence of this cassette, the plant’s internal defense signaling appears unable to recognize or neutralize the virus, leading to rapid disease progression.
Because Khufu can call and type these structural variants at scale, it provides a level of quantitative rigor that was previously impossible. Breeders can now quantify the "dosage" of resistance genes in their breeding lines, a task that was previously invisible to standard SNP-based diagnostic kits.
Official Perspectives: The Value of Genomic Clarity
The impact of this discovery extends beyond the laboratory. Dr. Jeremy Schmutz, a leader in the plant genomics field at HudsonAlpha, has emphasized that the transition to pangenomes is not just an incremental improvement—it is a fundamental change in how we view biological information.
"For too long, we have been viewing the world through a keyhole," researchers noted in their internal briefings. "By aligning everything to one reference, we were effectively ignoring the diversity that makes species resilient. Khufu allows us to see the entire landscape. The TSWV case study is the ‘proof of concept’ that validates this approach for other, equally complex agricultural challenges."
Industry stakeholders have lauded the shift, noting that the ability to select for structural variations in real-time allows for "targeted breeding." Instead of planting thousands of acres and hoping for a natural disease outbreak to test for resistance, breeders can now use molecular markers to identify the ideal genetic architecture in the seedling stage, shaving years off the development cycle of new, virus-resistant varieties.
Implications: The Future of Global Agriculture
The successful application of Khufu to the TSWV problem has profound implications for global food security.
1. Beyond TSWV: A Platform for Resistance
The researchers have hypothesized that this glutamate receptor locus may provide broad-spectrum viral resistance. If the structural variation identified here acts as a general defense mechanism, it could be leveraged to tackle a wider array of pathogens. The ability to "stack" these structural traits could lead to the development of "fortified" crops that are resilient to multiple viral strains simultaneously.
2. Economic Impact
For farmers, the economic stakes are astronomical. The billion-dollar losses associated with TSWV are not just financial figures; they represent the loss of livelihoods and the disruption of local food chains. By providing an actionable solution to a 20-year-old mystery, the Khufu approach directly addresses the financial instability that often plagues high-value crop production.
3. The End of the "Unsolvable" Problem
Perhaps the most significant implication is the change in the psychological and technical approach to breeding. For two decades, the resistance locus was considered "unsolvable" by conventional means. This breakthrough proves that many of the most persistent hurdles in crop science are not truly unsolvable; rather, they are simply invisible to the tools we have been using.
The Khufu approach is now being deployed for other traits, including drought tolerance, nutrient use efficiency, and yield architecture. By integrating low-pass sequencing with pangenome-guided detection, the research community is moving away from approximate marker associations toward precise, functional genomic insights.
Conclusion: A New Standard for Genomic Excellence
The story of the Khufu approach and its success with TSWV is more than just a technical victory; it is a fundamental reorientation of how we interact with the blueprint of life. By acknowledging the limitations of linear reference genomes and embracing the structural complexity of pangenomes, HudsonAlpha has provided a blueprint for the next century of plant breeding.
As we face the challenges of a growing global population and a changing climate, the need for high-precision, actionable genomic data has never been higher. The Khufu approach has turned a long-standing mystery into a roadmap for future success, ensuring that breeders can finally see—and utilize—the full spectrum of genetic variation. The era of the single reference is fading; the era of the pangenome has arrived.