For two decades, agricultural scientists and breeders grappled with a persistent, multi-billion-dollar adversary: the Tomato Spotted Wilt Virus (TSWV). Despite decades of effort, the genetic mechanism behind resistance remained elusive, obscured by the limitations of traditional sequencing technology. That changed recently when a team at the HudsonAlpha Institute for Biotechnology deployed "Khufu," a pioneering platform designed to overcome the constraints of conventional genomics. By moving beyond the single-reference genome, the team transformed a long-standing botanical mystery into a precise, actionable breeding strategy.
The Limitation of the Linear: Why Traditional Methods Failed
For years, the gold standard in genomics involved aligning short-read sequencing data against a single, static reference genome. While this method served the scientific community well for basic tasks, it possessed a fundamental flaw: reference bias. By forcing all genomic data to conform to one "standard" sequence, researchers often overlooked structural variations—complex, large-scale genetic rearrangements—that play a critical role in phenotypic expression.
In the case of TSWV, traditional genome-wide association studies (GWAS) repeatedly failed to pinpoint the exact causative variant. Researchers could identify general regions of interest, but they could never isolate the specific "switch" that conferred resistance. They were looking for a single-nucleotide polymorphism (SNP)—a small, single-letter change in the genetic code—when the true answer lay in a far more complex architectural arrangement.
The Khufu Innovation: Pangenomics at Scale
The Khufu approach represents a paradigm shift. Developed at HudsonAlpha, the platform is specifically engineered to maximize the efficiency and power of short-read, low-pass whole genome sequencing. Its secret weapon is "KhufuPAN," an add-on package that generates custom pangenome graphs.
Unlike a linear reference, a pangenome graph represents the collective genetic diversity of a population. It maps the genome as a network of variations, allowing researchers to see what is present in one individual compared to another, rather than just comparing everything to a single, potentially unrepresentative "template."
By applying this methodology to a large, segregating population of plants, the HudsonAlpha team was able to view the TSWV resistance locus through a high-resolution lens. Instead of searching for a needle in a haystack of SNPs, they were able to see the structure of the genome itself.
Chronology of a Discovery: From Mystery to Breakthrough
The journey to solving the TSWV puzzle was characterized by years of frustration followed by a rapid, high-impact resolution once the right tool was applied.
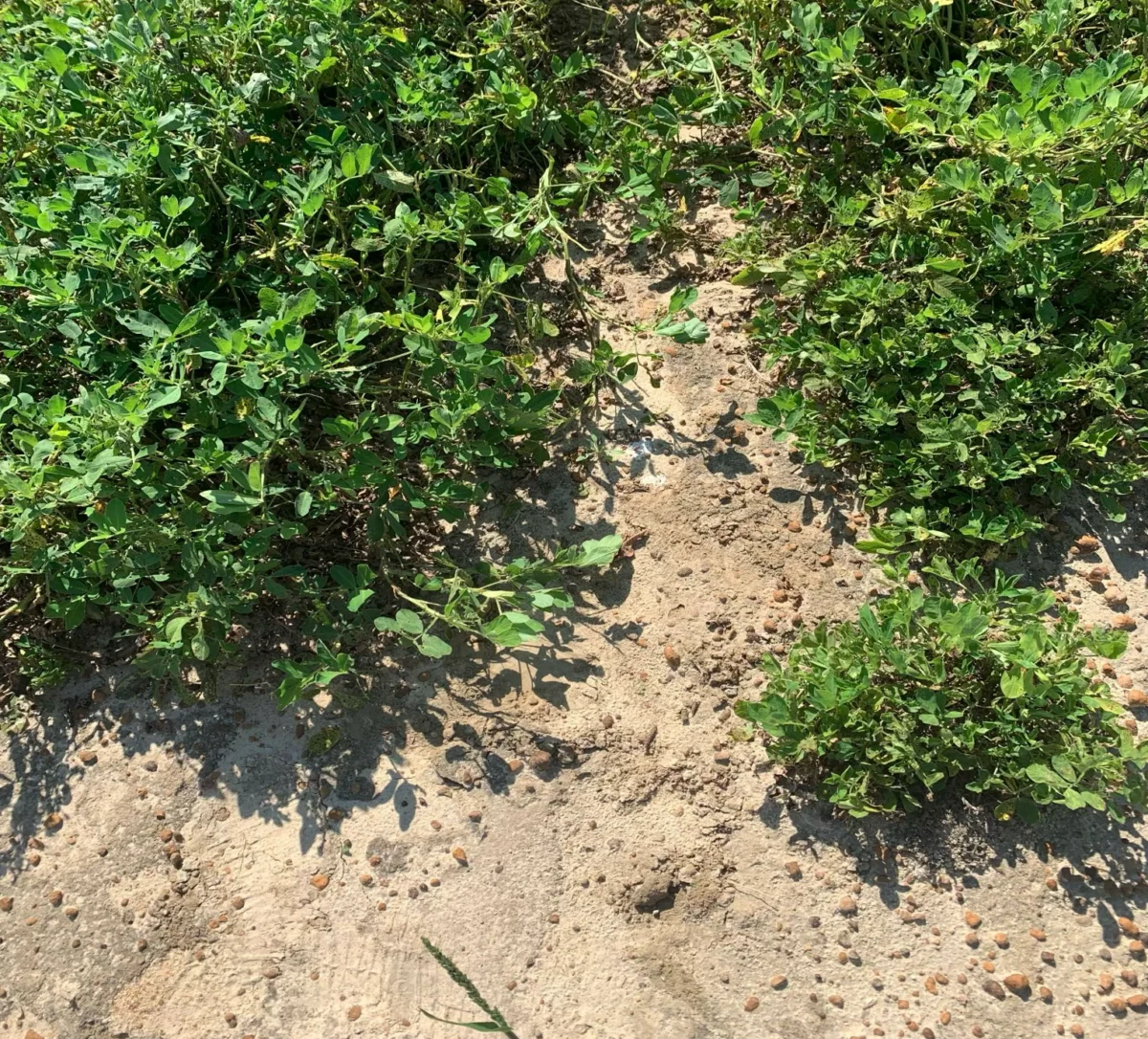
- 2004–2023: The Era of Ambiguity. For nearly 20 years, breeders observed varying levels of TSWV resistance in the field. Despite extensive phenotypic screening and traditional molecular marker analysis, the genetic basis remained elusive. Breeders relied on "proxy" markers that were often inconsistent across different breeding lines, leading to hit-or-miss outcomes.
- Early 2024: Deployment of the Khufu Platform. Researchers initiated a large-scale sequencing project using the Khufu workflow. By leveraging low-pass sequencing across thousands of individuals, the team gathered a massive, representative dataset of the population’s genomic landscape.
- Mid-2024: The Pangenome Reveal. Using KhufuPAN, the team constructed a pangenome graph that immediately highlighted a structural anomaly. They discovered a duplicated gene cassette containing four copies of a specific glutamate receptor gene.
- Late 2024: Phenotypic Validation. Correlating the copy number variation (CNV) with field data confirmed the discovery. Plants with four copies showed high resistance; those with fewer copies showed moderate resistance; and those lacking the duplication entirely were fully susceptible to the virus.
- Current Status: Implementation. Breeders are now integrating this precise "copy number" information directly into their marker-assisted selection programs, bypassing years of trial-and-error field testing.
Supporting Data: The Power of Copy Number Variation (CNV)
The TSWV study serves as a masterclass in why structural variants matter. The data revealed that resistance was not driven by a mutation within a gene, but by the dosage of a gene.
In genomic terms, this is a quantitative trait controlled by a structural variant. The Khufu system successfully called and typed these structural variants with a level of accuracy that standard pipelines—which often treat such duplications as mapping errors or "noise"—could not achieve. By treating these duplications as data points within a pangenome framework, Khufu provided a clear, quantitative correlation between the number of gene copies and the plant’s ability to survive viral infection.
This finding suggests that many "unsolved" breeding problems in other crops—such as drought tolerance, yield stability, or nutrient uptake—may also be driven by structural variants that remain hidden to traditional linear-based sequencing tools.
Official Perspectives: The Experts Weigh In
"We are no longer limited by the blind spots of the linear reference," notes the lead research team at HudsonAlpha. "Khufu allows us to see the full spectrum of variation. When we look at the genome as a graph, we see the evolutionary history and the structural complexity that makes each individual unique."
The sentiment among plant breeders who have adopted this approach is one of cautious optimism turning into excitement. "For years, we were flying blind," says one senior breeder involved in the validation trials. "We had an idea of the region, but we didn’t have the precision to make selection decisions with confidence. Now, we are selecting for the specific genomic configuration that guarantees resistance. It changes the economics of our entire breeding program."
Implications for Global Agriculture
The successful resolution of the TSWV resistance mystery has profound implications that extend far beyond a single plant species.
1. Accelerating Breeding Cycles
By identifying the exact structural cause of a trait, breeders can utilize "genomic selection" much earlier in the plant’s life cycle. Instead of waiting for a plant to mature and be exposed to environmental pressures—which may be inconsistent year-over-year—breeders can screen seedlings in the lab. This shortens the development cycle and reduces the cost of bringing improved varieties to market.
2. A New Standard for Complex Trait Discovery
The success of Khufu suggests that the scientific community should re-evaluate legacy data. Many projects that were deemed "unsuccessful" because they failed to find a causative SNP may contain the answers to vital agricultural questions, if only they were re-analyzed through the lens of a pangenome graph.
3. Addressing Broader Viral Resistance
The HudsonAlpha team is currently exploring whether the glutamate receptor gene cassette identified in this study might provide broad-spectrum resistance to other viruses beyond TSWV. If this locus acts as a master regulator of viral defense, it could become a cornerstone of genetic improvement programs for a wide variety of crops, potentially saving farmers billions in crop losses annually.
4. Democratizing Genomic Precision
Perhaps the most significant aspect of the Khufu approach is its efficiency. By optimizing for low-pass sequencing, the platform makes high-resolution, pangenome-scale analysis accessible to organizations that do not have the massive budgets required for long-read, high-coverage sequencing of every individual. This democratization of data means that smaller breeding programs and academic labs can now tackle the same complex genomic puzzles as global agricultural conglomerates.
Conclusion: A New Horizon in Genomics
The Khufu approach is more than just a new software package or a clever way to align reads. It represents a fundamental change in how we perceive the genome. For decades, the field of genetics was obsessed with finding the "single key" to unlock a trait. We now know that the genome is more like a complex, modular engine, where structural variations and gene copy numbers play as much of a role as the sequence of the letters themselves.
As the agricultural industry faces the mounting pressures of a growing population and a changing climate, the ability to rapidly identify and breed for resilience is no longer a luxury—it is a necessity. By turning a 20-year mystery into an actionable breeding strategy, the Khufu platform has proven that we don’t just need more data; we need more clarity. With this shift, the "unsolvable" problems of yesterday are becoming the standard solutions of tomorrow.