In the quest to secure global food supplies against the escalating pressures of climate change, researchers at the HudsonAlpha Institute for Biotechnology have achieved a monumental breakthrough. By mapping the complex genetic architecture of sorghum—a resilient, ancient cereal crop—scientists have moved beyond the limitations of a single "reference" genome, ushering in a new era of agricultural precision. This leap forward, described by lead researchers as the creation of a "genomic engine," promises to transform how we breed crops for extreme heat, drought, and pest resistance.
The Resilience of Sorghum: A Crop for a Warming World
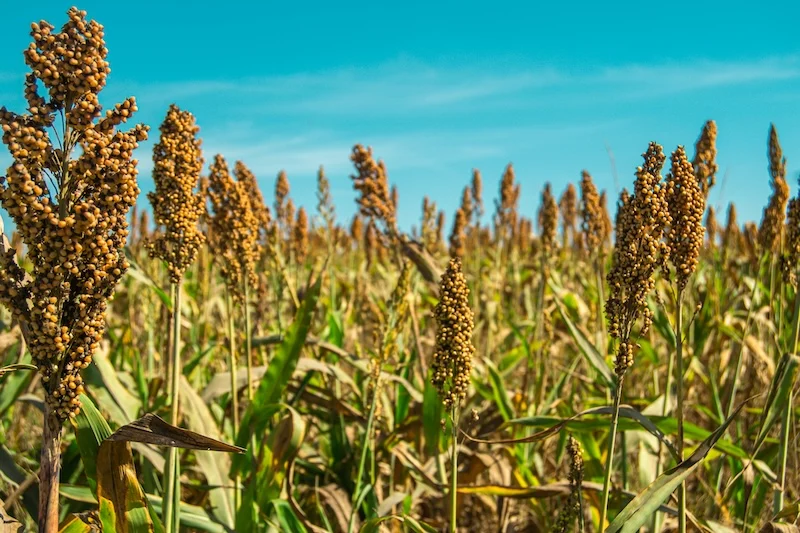
Sorghum (Sorghum bicolor) has long been celebrated as a miracle crop in arid and semi-arid regions of the world. Known for its incredible natural diversity, it thrives in environments where more sensitive staples, such as maize or wheat, often fail. From the sub-Saharan plains to the drylands of North America, sorghum serves as a critical source of food, feed, and biofuel.
Despite its robust nature, the very diversity that makes sorghum a survivalist has historically served as a hurdle for agricultural science. "Sorghum has incredible natural diversity that allows it to grow in places where other crops fail," explains John Lovell, PhD, a HudsonAlpha Research Faculty Investigator and the lead researcher on this project. "However, that same diversity has historically made it difficult to breed sorghum with precision. Our lab focused on building the ‘engine’ for this project, creating the genomic tools and maps that allow other scientists to finally see the whole picture."
A Chronology of Genomic Innovation
To understand the magnitude of this achievement, one must look back at the state of genomic science over the past decade.
2011: The Era of the "Reference" Genome
Since 2011, the scientific community has relied on a single "reference" genome to represent the entirety of the sorghum species. This approach treated one specific variety as the template for all others, an outdated methodology akin to using a single map to navigate an entire continent.
2011–2023: The Limitations of "One-Size-Fits-All"
For over a decade, plant breeders and geneticists struggled with the "reference bias." The single-genome approach often ignored large sections of DNA—known as structural variants—that define why one variety of sorghum can survive record-breaking heat while another succumbs to a common pest. These "hidden" sequences were essentially invisible to researchers using the 2011 model.
2024: The Pangenome Breakthrough
The team at HudsonAlpha, working in collaboration with a global consortium of researchers, moved to develop a "pangenome." Unlike a single reference, a pangenome captures the entire genetic diversity of a species by mapping multiple varieties simultaneously. This transition from a linear reference to a multidimensional graph-based map marks the most significant advancement in sorghum research in recent history.
Supporting Data: Dissecting the "Engine"
The core of the HudsonAlpha achievement lies not just in the discovery of specific genes, but in the creation of a scalable infrastructure that can be utilized by labs worldwide. By developing advanced genomic tools and maps, the team has effectively digitized the biological diversity of sorghum.
Key Discoveries Enabled by the Infrastructure
The study highlighted several high-impact breakthroughs that were made possible only through the new pangenome model:
- Seed Shattering Identification: Researchers successfully identified a specific sequence insertion responsible for "seed shattering"—a trait where seeds fall from the plant prematurely, causing significant yield loss.
- Tracing Gene Flow: By analyzing the pangenome, the team was able to trace how specific genes have moved through modern breeding programs, allowing for a better understanding of how current varieties have been shaped by human selection.
- Structural Variant Mapping: The new resources allow scientists to view structural variations—large-scale rearrangements of DNA—that were previously invisible, which are often the primary drivers of environmental adaptation.
Official Responses and Expert Perspectives
The implications of this research are resonating throughout the agricultural and biotechnological communities. Jeremy Schmutz, HudsonAlpha Faculty Investigator and co-director of the Genome Sequencing Center (GSC), emphasized that the utility of these tools lies in their accessibility and versatility.
"These tools are far-reaching because each researcher can use them for their own specific needs," Schmutz stated. "Whether a scientist is looking for resistance to the parasitic Striga weed or better drought tolerance, they can now query an interval of interest, dissect it, and dive deep into the pangenome variation. It transforms foundational biology into actionable breeding decisions."
For breeders, the shift is profound. Historically, identifying the genetic basis for a trait like drought resistance could take years of trial and error. With the pangenome "engine," researchers can pinpoint the exact gene responsible for a trait in a matter of weeks, accelerating the development of crops that can withstand the climate conditions of the next century.
Implications: Building a Resilient Future
The global food system is currently facing a "perfect storm" of challenges: a growing population, dwindling water resources, and an increasingly volatile climate. The ability to "read" the genetic potential of sorghum with this level of accuracy is a cornerstone of future-proofing agriculture.
Transforming Foundational Biology into Actionable Breeding
The transition from basic genomic research to "actionable breeding decisions" is the ultimate goal of the HudsonAlpha project. By providing an open-access library of sorghum’s genetic diversity, the institute is democratizing the ability to improve crops.
- Climate Adaptation: Breeders can now select for varieties that possess the specific structural variants required to thrive in high-heat or low-moisture conditions.
- Pest and Disease Resistance: With the ability to query specific genetic intervals, scientists can target the genes that trigger immune responses against weeds like Striga and various insect populations, potentially reducing the need for chemical pesticides.
- Yield Stability: By eliminating detrimental traits like seed shattering and enhancing desirable traits through informed selection, the consistency of the global sorghum harvest can be drastically improved.
The Broader Impact on Global Agriculture
The work conducted by the HudsonAlpha team serves as a blueprint for other cereal crops. If the pangenome approach can work for sorghum, it can be applied to maize, rice, and wheat. As researchers continue to build upon this "global library" of genetic information, the scientific community is moving closer to a state where agricultural production is no longer at the mercy of environmental unpredictability.
Conclusion: The Next Horizon
The genomic tools developed at HudsonAlpha are more than just a scientific accomplishment; they are a vital asset for global food security. By moving past the constraints of a single reference genome, researchers have unlocked the hidden potential within sorghum’s DNA.
As the world continues to navigate the complexities of a changing climate, the "genomic engine" created by Dr. Lovell, Jeremy Schmutz, and their team will undoubtedly become a foundational component of 21st-century agriculture. It provides the roadmap needed to navigate the challenges of tomorrow, ensuring that crops like sorghum can continue to feed a hungry planet in the most difficult of circumstances.
The era of precise, rapid, and inclusive crop improvement has arrived. Through the power of the pangenome, the hidden secrets of plant evolution are finally being translated into the solutions we need to survive and thrive.