Tübingen, Germany – April 12, 2026 – In a significant advancement for cancer research and precision medicine, a team of Tunisian scientists has unveiled "OncoSolidDB," a comprehensive and meticulously curated bioinformatics database focused on ligands targeting solid tumors. Published in the prestigious journal Cancers, this novel resource promises to streamline drug discovery, accelerate the repurposing of existing therapies, and foster the rational design of next-generation treatments for a wide spectrum of solid malignancies.
The Critical Need for a Specialized Oncology Ligand Database
Cancer continues to be a formidable global health challenge, with projections indicating a dramatic rise in new cases and mortality rates. Solid tumors, encompassing common cancers such as breast, lung, colorectal, and prostate, represent the vast majority of diagnoses and present unique therapeutic hurdles due to their heterogeneity and the potential for drug resistance. While targeted therapies have transformed oncology, the information regarding the ligands that mediate these treatments is often fragmented across numerous databases, hindering efficient research.

"Researchers have been facing a significant challenge in accessing structured, oncology-specific data on therapeutic ligands in a consolidated manner," explains Dr. Oussema Khamessi, lead author of the study and a researcher at the Laboratory of Bioinformatics, Biomathematics and Biostatistics (BIMS-LAB) at Institut Pasteur de Tunis. "This fragmentation makes it difficult to conduct comprehensive analyses, perform comparative pharmacology, and ultimately, to accelerate the development of more effective treatments."
To address this critical gap, Khamessi and his colleagues developed OncoSolidDB, a freely accessible, open-access database that consolidates and harmonizes data on ligands specifically targeting solid tumors.

Unveiling OncoSolidDB: A Curated Treasure Trove of Ligand Data
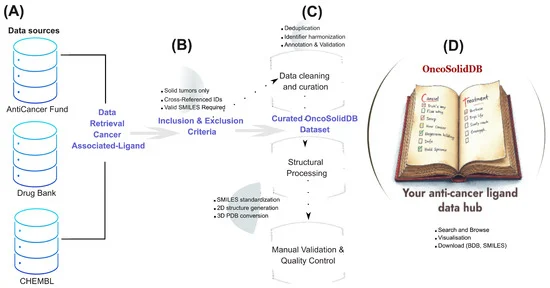
The newly launched OncoSolidDB is the culmination of extensive data collection and rigorous curation efforts. The database integrates information from authoritative sources such as ChEMBL, DrugBank, and the Anti-Cancer Fund. Each ligand entry is meticulously annotated with standardized identifiers, chemical structures (including SMILES strings and 2D images), pharmacological details, and crucial regulatory approval history. A key feature is the inclusion of downloadable protein structural files (PDB format), which are essential for advanced structural bioinformatics analyses.
Currently, OncoSolidDB houses data on 243 ligands that have been linked to 15 major solid cancer types. These include commonly diagnosed cancers like breast, lung, colorectal, and prostate, as well as others such as ovarian, cervical, bladder, esophageal, gastric, head and neck, thyroid, pancreatic, renal, and liver (Hepatocellular Carcinoma).

Chronology of Development and Data Integration
The journey to OncoSolidDB began with a systematic data collection phase. Researchers queried established databases like DrugBank and ChEMBL, specifically searching for compounds associated with solid tumor indications. The Anti-Cancer Fund database served as an additional crucial source.
Following initial data retrieval, a rigorous multi-step curation and standardization pipeline was implemented. This process involved:

- Eligibility Filtering: Strict inclusion and exclusion criteria were applied to ensure only high-confidence, oncology-relevant ligands were retained. Ligands were required to have validated cancer indications, valid chemical structures, and cross-referenced identifiers. Compounds exclusively linked to hematological malignancies, those lacking structural data, or duplicates were excluded.
- Data Curation and Standardization: Manual validation of cancer indications, correction of inconsistencies in ligand names, and standardization of SMILES representations were performed. Cross-referencing across databases enriched each entry with consistent annotations.
- Structural Processing: For computational compatibility, ligands were converted into SMILES strings, 2D images, and downloadable 3D PDB files using robust bioinformatics tools such as RDKit and Open Babel.
- Web Interface Development: A user-friendly web interface was built using Flask, HTML5, and CSS3, enabling intuitive browsing, searching, and data retrieval.
The temporal coverage of the data spans from 1953 to 2025, offering researchers a valuable historical perspective on the evolution of oncology drug development.
Supporting Data: Unveiling Trends and Overlaps
The analysis of OncoSolidDB’s content has revealed several significant trends and insights:

- Cancer Type Distribution: Lung cancer (18.9%) and breast cancer (16.8%) feature the highest number of associated ligands, likely reflecting their prevalence and the extensive research dedicated to these diseases. Ovarian cancer and melanoma follow, with prostate and pancreatic cancers also showing substantial representation. Conversely, gastric and head and neck cancers are currently less represented, indicating potential areas for future data expansion.
- Temporal Trends: The database shows a marked increase in ligand approvals from the 1990s onward, with a significant acceleration observed between 2015 and 2025. This pattern underscores the shift towards targeted therapies and precision medicine in recent decades.
- Chemical Complexity: Ligand complexity, as proxied by SMILES string length, varies widely, with most ligands falling within a moderate range of complexity.
- Ligand Overlap Analysis: An innovative UpSet plot analysis revealed that while many ligands are specific to a single cancer type, a notable number are shared across multiple solid tumors. This overlap highlights common underlying oncogenic mechanisms and presents significant opportunities for drug repurposing. For example, ligands targeting HER2 (like trastuzumab) and those inhibiting angiogenesis (like bevacizumab) are found across several cancer types, demonstrating their broad therapeutic potential.
Official Responses and Broader Implications
The development of OncoSolidDB has been met with enthusiasm from the scientific community. "This database is a game-changer for translational oncology research," commented Dr. Kais Ghedira, a senior author and head of the BIMS-LAB. "By providing a centralized, cancer-specific resource with integrated structural data, we are empowering researchers to conduct more sophisticated analyses and to accelerate the discovery of novel therapeutic strategies."
The implications of OncoSolidDB are far-reaching:

- Drug Discovery and Development: Researchers can now efficiently identify potential drug candidates for specific solid tumors, perform virtual screening, and conduct molecular docking studies with greater ease.
- Drug Repurposing: The analysis of ligand overlap across cancer types directly supports the identification of existing drugs that could be repurposed for new indications, potentially reducing development time and costs.
- Precision Medicine: By offering detailed ligand information tailored to specific cancer types, OncoSolidDB facilitates the design of personalized treatment plans based on the molecular profile of a patient’s tumor.
- Educational Resource: The database serves as an invaluable educational tool for students and researchers entering the field of cancer bioinformatics and drug discovery.
Future Directions and Vision
The research team is committed to the continuous development and expansion of OncoSolidDB. Future plans include incorporating additional investigational compounds, ligands currently in clinical trials, and newly approved therapeutics. Furthermore, the integration of ligand-target interaction data and protein structural information will enhance the database’s utility for complex structural bioinformatics analyses.
"Our vision is to make OncoSolidDB a dynamic and evolving resource that remains at the forefront of cancer research," stated Dr. Ghedira. "We aim to leverage artificial intelligence and high-performance computing to generate AI-ready datasets, further supporting advanced machine learning and deep learning approaches for predicting ligand-receptor interactions and optimizing drug design."

OncoSolidDB is publicly accessible via its dedicated web portal, https://liganddb.pythonanywhere.com/solid_cancer, and also through the Bioinformatic Research Portal "Einstein" at http://196.203.66.81/. This initiative represents a significant stride towards a more integrated and efficient approach to tackling the complex challenge of solid tumors, ultimately promising to improve patient outcomes worldwide.