The global crisis of antimicrobial resistance (AMR) has long demanded a shift in how we identify new therapeutic agents. Traditional drug discovery pipelines are frequently plagued by high attrition rates, where promising compounds fail not due to a lack of target affinity, but because of poor "developability"—the complex interplay of solubility, permeability, safety, and stability. A new study published in the journal Pharmaceuticals (Vol. 19, 744) introduces a novel, computation-assisted framework designed to bridge this gap: the Fivefold Maximum Drug-Likeness (5F-MDL) strategy.
By integrating deep learning with a multidimensional screening approach, researchers from Tongren University and Yangtze University have demonstrated a potential roadmap for prioritizing antibacterial candidates against Escherichia coli (E. coli), one of the most persistent and dangerous Gram-negative pathogens.

The Core Concept: Moving Beyond Simple Affinity
Drug discovery has historically leaned on high-throughput screening (HTS) or structure-based rational design. While HTS can process millions of molecules, it is notoriously inefficient, often yielding "false positives" that lack the necessary pharmacokinetic properties to function in a complex biological environment. Rational design, while precise, is often hampered by limited structural data.
The 5F-MDL strategy, led by researchers Haoyu Zhu, Lu Xu, and Wei Shi, addresses these limitations by shifting the focus from a single-objective search to a holistic, multidimensional evaluation. "Fivefold" refers to the five critical dimensions of drug developability: physicochemical properties, pharmacokinetics, efficacy, safety, and stability.

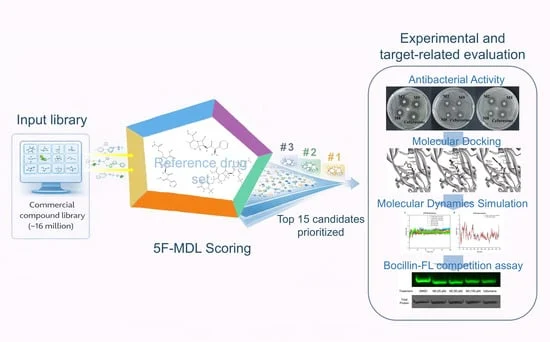
Instead of relying on a single scalar value like a docking score, the team developed an ensemble of 33 deep learning submodels. These models generate a 33-dimensional "property spectrum" for each molecule, allowing researchers to assess how closely a candidate aligns with the known profiles of successful, clinically approved cephalosporin antibiotics.
Chronology of the Study
The research, which culminated in the May 2026 publication, followed a rigorous multi-stage timeline:

- March 29, 2026: The manuscript was officially submitted for peer review.
- April 2026: The study underwent a comprehensive revision process, refining the computational models and clarifying the experimental validation protocols.
- May 5, 2026: The research was officially accepted, following successful demonstration of its methodology.
- May 8, 2026: The article was published, presenting the 5F-MDL paradigm to the scientific community.
The experimental validation involved screening approximately 16 million commercially available molecules. From this massive library, the team utilized their 5F-MDL scoring system to prioritize 15 top-ranked candidates for direct laboratory testing against E. coli ATCC 25922.
Supporting Data and Experimental Findings
The validation phase was designed to stress-test the computational predictions. Of the 15 candidates selected, three compounds—designated M2, M8, and M9—demonstrated measurable antibacterial activity in disk diffusion assays.

The Rise of Candidate M2
Among the three active compounds, M2 emerged as the most promising. In broth microdilution assays, M2 displayed an MIC (Minimum Inhibitory Concentration) of 25.6 µg/mL and an MBC (Minimum Bactericidal Concentration) of 51.2 µg/mL. While these results show that M2 is less potent than the reference antibiotic cefuroxime (which exhibited an MIC of 3.2 µg/mL), the successful translation of a computer-prioritized hit into a functional antibacterial molecule serves as a proof-of-concept for the 5F-MDL framework.
Mechanistic Insights
To understand how these molecules interact with their target, the researchers turned to molecular docking and dynamics simulations, focusing on Penicillin-Binding Protein 2 (PBP2). The simulations suggested that M2 maintains a stable, noncovalent interaction pattern within the PBP2 active pocket. Further biochemical validation via a Bocillin-FL competition assay confirmed that M2 could effectively interfere with the fluorescent probe labeling of PBP2 in a concentration-dependent manner, providing strong, though preliminary, evidence that the compound targets the cell wall biosynthesis pathway.

Professional and Official Perspectives
The authors of the study, including lead researchers from Tongren University, have been careful to temper the excitement surrounding these results. In their official discussion, they state: "M2 should be regarded as a preliminary antibacterial hit for further optimization rather than as a validated lead compound."
The research team acknowledges that the 5F-MDL framework is in its infancy. They emphasize that while their deep learning models showed strong internal consistency—achieving high coefficients of determination (R²) across the 33 property-specific submodels—these results do not yet guarantee the same success rate when applied to resistant clinical isolates. The study explicitly invites the broader scientific community to perform benchmarking against established prioritization methods to determine the true comparative advantage of this multidimensional approach.

Implications for Future Antibiotic Discovery
The 5F-MDL strategy represents a significant evolution in computer-aided drug design (CADD). By forcing a "developability-aware" perspective, the framework essentially acts as a filter that eliminates candidates prone to failure long before they reach the expensive and time-consuming stage of wet-lab testing.
Addressing the Gram-Negative Bottleneck
Gram-negative bacteria like E. coli are notoriously difficult to treat due to their complex outer membranes, which act as a physical barrier to many drugs. The 5F-MDL approach is particularly relevant here; because it considers properties like permeability and intracellular accumulation, it can identify molecules that possess the "balanced" profile required to breach the Gram-negative barrier.

Next Steps
The research team has identified several pathways for future improvement:
- Broader Validation: Future work will focus on testing optimized candidates against drug-resistant clinical strains, such as carbapenem-resistant Enterobacteriaceae (CRE).
- Model Refinement: There is a clear need to integrate more Gram-negative-specific parameters, such as efflux pump susceptibility and membrane permeability metrics, into the 33-dimensional property spectrum.
- Benchmarking: Formal head-to-head comparisons against conventional methods like QED (Quantitative Estimate of Drug-likeness) are necessary to prove that 5F-MDL offers a superior return on investment for pharmaceutical R&D.
In conclusion, the 5F-MDL strategy is a bold step toward a more predictive, resource-efficient future in drug discovery. While the path from a computational hit like M2 to a clinical antibiotic is long and arduous, the ability to prioritize candidates with balanced, developability-aware profiles provides a necessary tool in the ongoing battle against the global threat of antibiotic resistance. The study provides a compelling case that in the era of artificial intelligence, the best way to find a cure is to consider the whole molecule, not just the target binding site.